ICLR LMRL Workshop, 2026 Spotlight
Incorporating contextual information into KGWAS for interpretable GWAS discovery
1University of Michigan
2gRED, Genentech
3Stanford University
§Work conducted during an internship at Genentech
†Correspondence: {yao.heming, hoeckendorf.burkhard, richmond.david} AT gene DOT com
Abstract
Genome-Wide Association Studies (GWAS) identify associations between genetic variants and disease; however, moving beyond associations to causal mechanisms is critical for therapeutic target prioritization. The recently proposed Knowledge Graph GWAS (KGWAS) framework addresses this challenge by linking genetic variants to downstream gene-gene interactions via a knowledge graph (KG), thereby improving detection power and providing mechanistic insights. However, the original KGWAS implementation relies on a large general-purpose KG, which can introduce spurious correlations. We hypothesize that cell-type specific KGs from disease-relevant cell types will better support disease mechanism discovery. Here, we show that the general-purpose KG in KGWAS can be substantially pruned with no loss of statistical power on downstream tasks, and that performance further improves by incorporating gene–gene relationships derived from perturb-seq data. Importantly, using a sparse, context-specific KG from direct perturb-seq evidence yields more consistent and biologically robust disease-critical networks.
Workflow
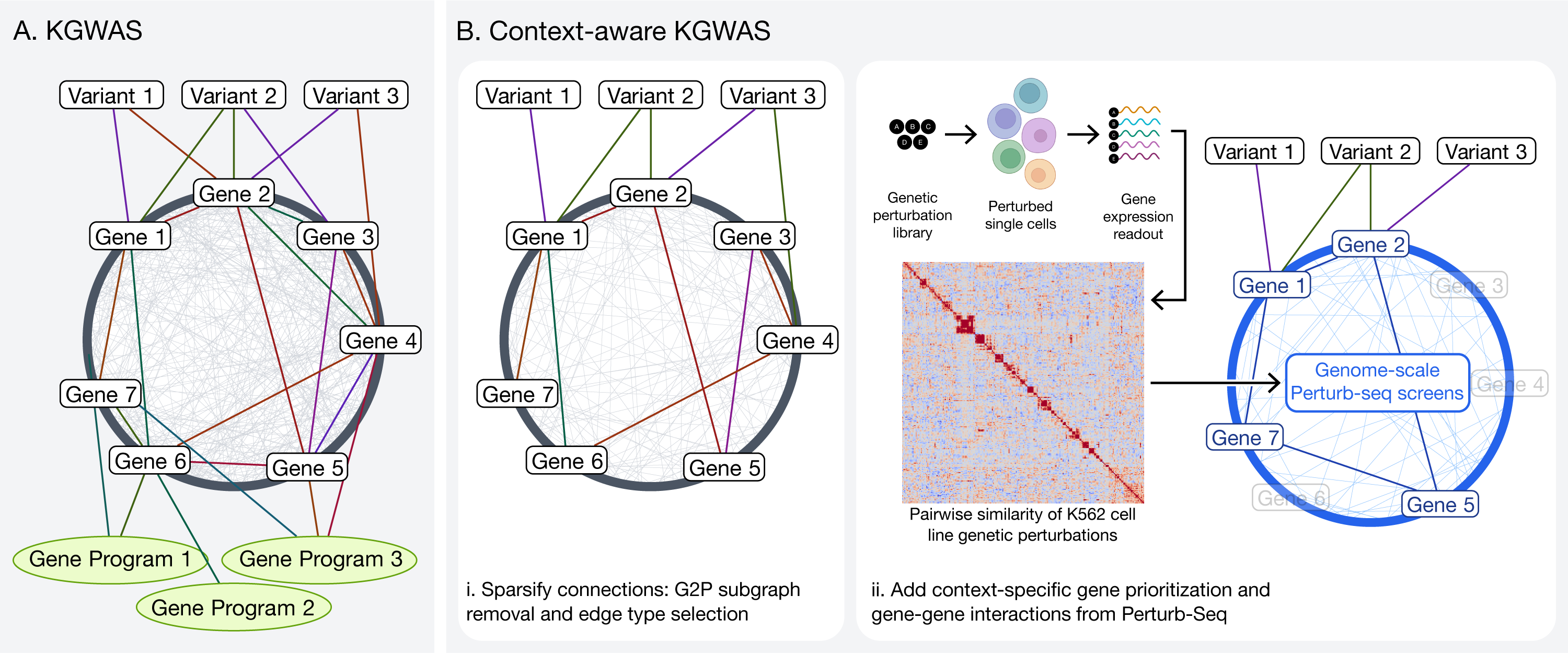
A. The original KG in KGWAS consisting of variant-gene, gene-gene and gene-program edges; B. Our extension to KGWAS: (left) removing gene-program edges, and sparsifying remaining connections; (right) replacing gene-gene edges with contextually relevant relationships derived from Perturb-seq.
Context-aware KGWAS improves hit detection while reducing graph size
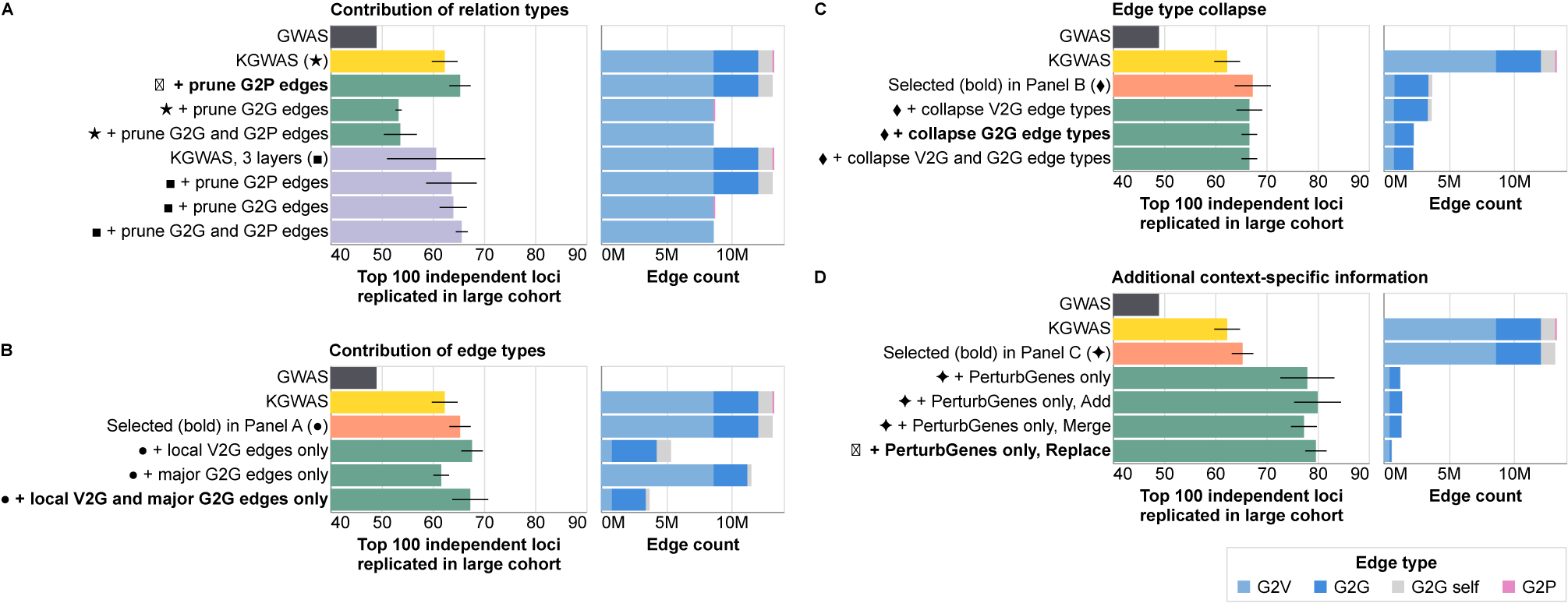
Contributions of different nodes and edge types in the KGWAS knowledge graph using a sample size of 10,000. Reported metrics are the total number of recalled independent loci summed across three selected traits. Standard deviations are computed across three training runs with different random seeds. In each subplot, bold indicates the model selected as the baseline in the subsequent panel. (A) Ablation of different relation types; (B) Ablation of V2G and G2G edge types; (C) Collapsing V2G and G2G edge types; (D) Adding context-specific information.
Context-aware KGWAS improves interpretability of disease critical networks
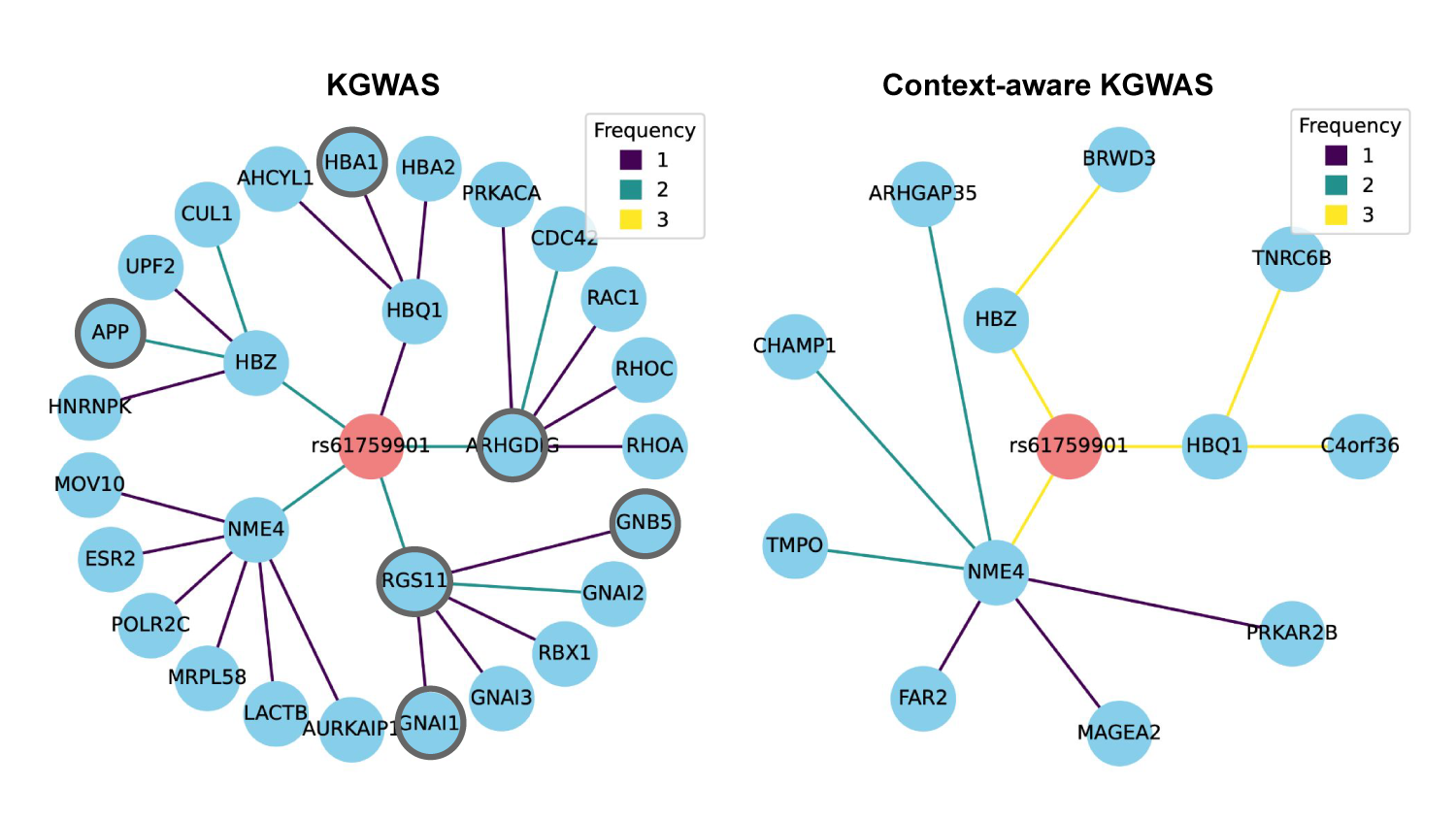
Consistency of disease critical networks for the rs61759901 variant in KGWAS (left) and context- aware KGWAS (right). Each plot aggregate nodes and edges from three seeded models trained on the MCH trait. Genes that are not significant in the K562 cell line (n=6) are shown with a gray border.
Bibtex
@inproceedings{
jiang2026incorporating,
title={Incorporating contextual information into {KGWAS} for interpretable {GWAS} discovery},
author={Cheng Jiang and Brady Ryan and Megan Crow and Kipper Fletez-Brant and Kashish Doshi and Sandra Melo and Kexin Huang and Burkhard Hoeckendorf and Heming Yao and David Richmond},
booktitle={Learning Meaningful Representations of Life (LMRL) Workshop at ICLR 2026},
year={2026},
}